This research tackles antibiotic resistance by developing nano-scale microfluidic cultures that isolate and study previously unculturable bacteria. By screening rare microbes and directly testing their antimicrobial activity, the platform accelerates discovery of new antibiotics, offering a powerful tool against drug-resistant superbugs.
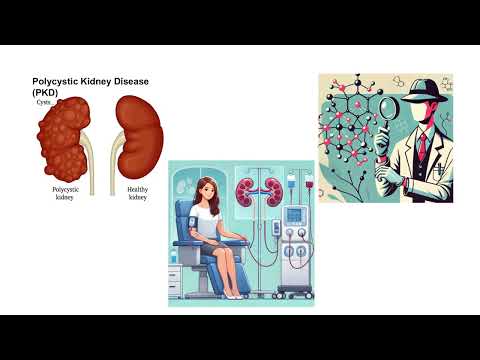
The speaker studies polycystic kidney disease by identifying missing or damaged proteins that destabilize the kidney’s filtration network. Using BioID and mass spectrometry, they map healthy versus diseased protein interactions to pinpoint weak spots. This work aims to enable targeted therapies and personalised treatments for PKD patients.
This research provides the first-ever map of the honeybee gut protein interactome to understand how the parasite Nosema disrupts bee health. By isolating gut protein interactions and identifying them via mass spectrometry and computational analysis, the project uncovers how infection alters essential networks, paving the way for targeted, safer treatments for honeybee disease.
This research develops protein-based forensic tools to detect meat adulteration in processed foods. By designing species-specific protein biomarkers and using mass spectrometry, the method identifies hidden pork, beef, chicken, or lamb in mixed meat products. The approach supports food safety, religious dietary compliance, allergy protection, and government efforts to combat food fraud.