This research develops targeted radiopharmaceutical therapies for HER2-positive cancers. By attaching radioactive isotopes to trastuzumab, treatment delivers precise radiation to cancer cells, overcoming drug resistance. The work includes creating practical drug kits and aims to improve cancer outcomes by replacing non-specific therapies with highly accurate, targeted interventions.
This research applies machine learning to genetic data to distinguish harmless DNA variations from cancer-causing mutations. By treating DNA like a crime scene, the model learns to identify which genetic changes truly drive breast cancer risk, supporting more accurate diagnosis and informed clinical decision-making.
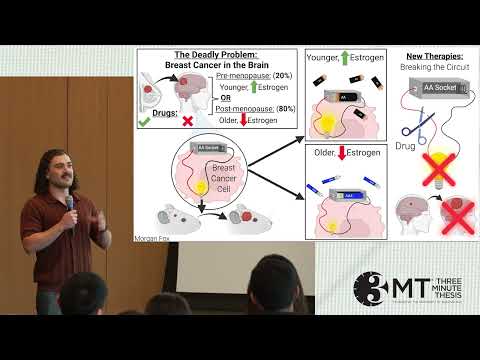
Breast cancer most often kills by spreading to the brain, where hormone therapies fail. This research reveals a signaling pathway that drives tumor growth in both pre- and postmenopausal settings. Identifying alternative activators of this pathway opens new therapeutic opportunities for deadly brain metastases.
This research improves preclinical testing of SERDs (selective estrogen receptor degraders) for estrogen receptor–positive breast cancer by modeling the tumor environment and treatment resistance. By co-culturing cancer and fat cells and applying single-cell RNA sequencing, it identifies resistance mechanisms to support more effective drug development for patients.
This research designs bespoke, lightweight 3D-printed breast prostheses for women after mastectomy. Using body scanning and personalised materials, the prostheses improve comfort, symmetry, and fit compared to standard silicone forms. The project aims to help women move from surviving breast cancer to thriving with confidence and comfort.
This research examines how macrophages shift between tumor-fighting and tumor-supporting roles in breast cancer. By identifying signals in the tumor microenvironment and engineering molecular cues to promote tumor-destroying behavior, the work aims to reprogram immune responses and improve therapeutic outcomes for breast cancer patients.
My research investigates collagen-binding receptors on breast cancer cells as potential biomarkers to distinguish harmless early-stage tumours from aggressive ones. Using genetically matched 3D cancer models, the project identifies how receptor activity affects invasion and collagen organization, aiming to reduce overtreatment and support clearer clinical decisions for early breast cancer patients.
The talk explains how drug discovery struggles with the enormous size of chemical space, where only a few molecules become effective medicines. Using miniaturized chemical libraries and off-rate screening, the researcher accelerates structure–activity relationships (SAR) mapping without purification. This approach has already produced promising breast-cancer drug candidates and could dramatically reduce drug-development costs.